This was a collaborative project between our lab and Dr. Gene Robinson’s behavioral neuroscience lab and involves the application of systems approaches to predict the effect of a dynamic process like behavior on gene expression.
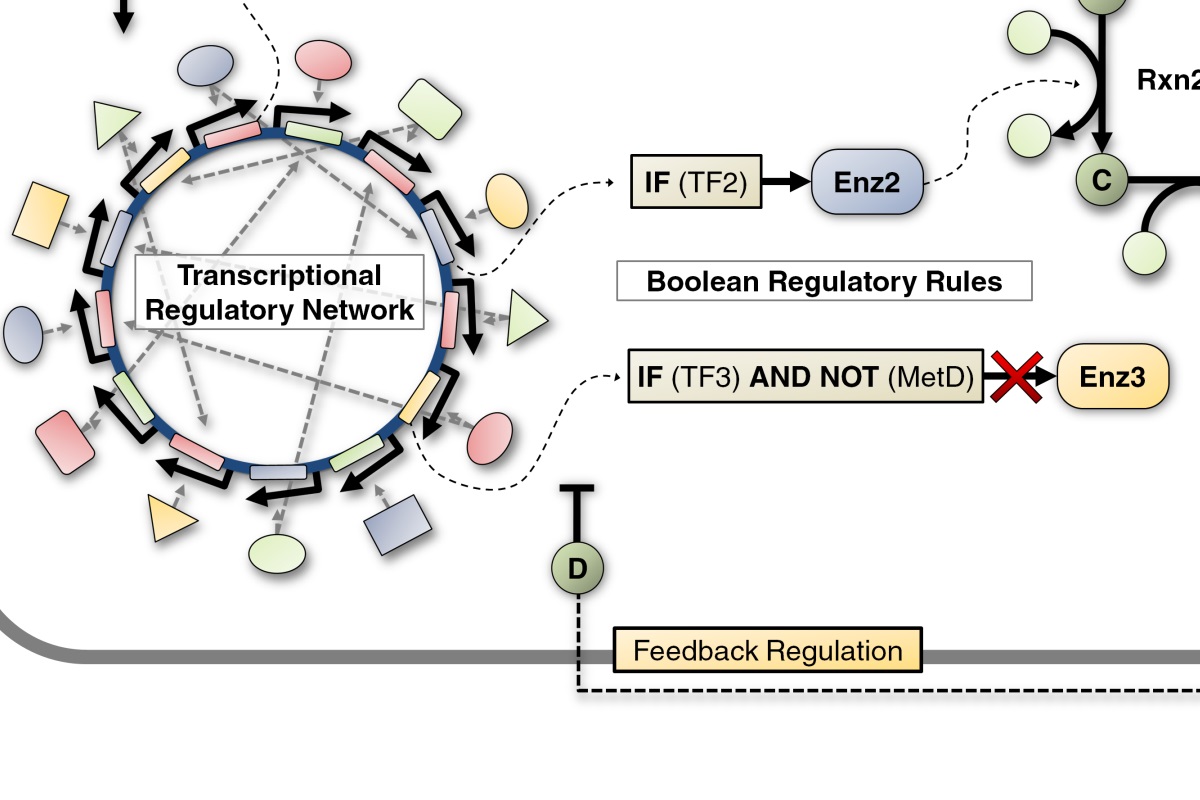
Using a brain transcriptome data set (generated in Dr. Robinson’s lab) unparalleled in size and scope (over 800 individual honeybees sampled in almost 50 behavioral states) we succeeded in reconstructing a Transcriptional Regulatory Network (TRN) that showed remarkably high accuracy in quantitatively predicting brain gene expression. This TRN encompasses thousands of genes and we found specific transcription factors that are central actors in regulating behavior.
TRN reconstruction was performed with a new combination of two well-known algorithms; the ASTRIX (Analyzing Subsets of Transcriptional Regulators Influencing eXpression) algorithm generates a network of high-confidence TF-gene interactions using ARACNE (Accurate Reconstruction of Cellular Networks), and leverages these interactions to predict expression in new conditions using LARS (Least Angle Regression). Our results show that despite the daunting anatomical and physiological complexity of the brain, simple linear connections between transcription factors and their putative target genes are a surprisingly prominent feature of brain neurogenomic states that underlie naturally occurring behavior.
Data and Software File(s):
TRN in Excel format


 hood-price.isbscience.org/research/honeybee-transcriptional-regulatory-network/
hood-price.isbscience.org/research/honeybee-transcriptional-regulatory-network/